Abstract
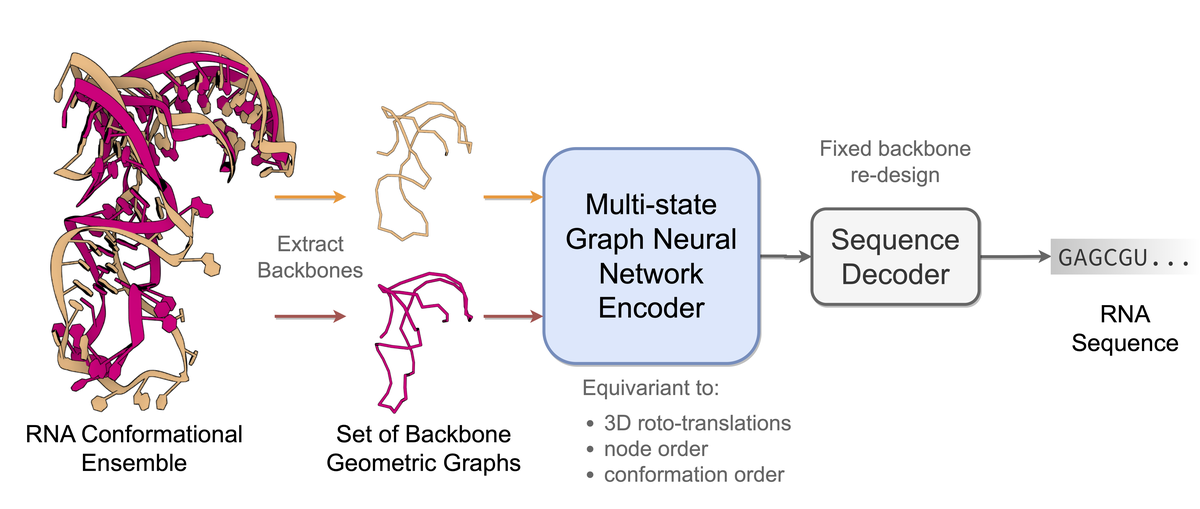
omputational RNA design tasks are often posed as inverse problems, where sequences are designed based on adopting a single desired secondary structure without considering 3D conformational diversity. We introduce gRNAde, a geometric RNA design pipeline operating on 3D RNA backbones to design sequences that explicitly account for structure and dynamics. gRNAde uses a multi-state Graph Neural Network and autoregressive decoding to generates candidate RNA sequences conditioned on one or more 3D backbone structures where the identities of the bases are unknown. On a single-state fixed backbone re-design benchmark of 14 RNA structures from the PDB identified by Das et al. (2010), gRNAde obtains higher native sequence recovery rates (56% on average) compared to Rosetta (45% on average), taking under a second to produce designs compared to the reported hours for Rosetta. We further demonstrate the utility of gRNAde on a new benchmark of multi-state design for structurally flexible RNAs, as well as zero-shot ranking of mutational fitness landscapes in a retrospective analysis of a recent ribozyme. Experimental wet lab validation on 10 different structured RNA backbones finds that gRNAde has a success rate of 50% at designing pseudoknotted RNA structures, a significant advance over 35% for Rosetta.